19th Century New England Vampire Undergoes 21st Century DNA Analysis
Cutting-Edge Laboratory and Bioinformatics Techniques Were Evaluated to Reveal What He Looked Like and Confirm Who He Was
RESTON, VIRGINIA, UNITED STATES, October 30, 2022 /EINPresswire.com/ -- Tales of the undead consuming the blood of living beings have been around for centuries. Before scientific and clinical knowledge were used to explain infectious diseases and medical disorders, communities hit with epidemics turned to folklore for explanations. They often blamed vampirism for the change in appearance, erratic behavior and deaths of their friends and family who actually suffered from conditions such as porphyria, pellagra, rabies and tuberculosis (TB).
At the International Symposium on Human Identification (ISHI) conference, being held in Washington, D.C. this week, Parabon NanoLabs and the Armed Forces DNA Identification Laboratory (AFDIL, a branch of the Armed Forces Medical Examiner System)* will unveil results from the latest advanced laboratory and bioinformatic DNA analyses performed on early 19th-century unidentified remains of a man known as JB55 because of the markings on his grave. The remains, discovered in 1990 in Griswold, CT, were determined to be those of a middle-aged man who suffered from tuberculosis. It is speculated that he was later disinterred and reburied because his limbs had been placed atop his chest in an X in a skull-and-crossbones configuration — a burial practice used to prevent purported vampires from rising from the grave to feed upon the living.
In 2019 AFDIL performed a Y-STR analysis that suggested a possible surname of “Barber". A search of historical records led to an obituary for another individual buried in the same cemetery that mentioned a man named John Barber, but no other records were found for him(1).
The DNA work in this study builds on a previous collaboration between Parabon and AFDIL that targeted ~95,000 single nucleotide polymorphisms (SNPs) for direct kinship comparisons using degraded DNA(2). Answering questions like: Was “JB55” in fact John Barber as previously speculated in 2019? And, what did JB55 look like? require ~10x as much data, and the purpose of the current study was to determine what laboratory and bioinformatics methods are best able to create a genome-wide SNP data profile to answer those questions.
Old bone samples are challenging for DNA analysis both because the human DNA can be highly degraded and because a large majority of the sample may consist of non-human DNA from the environment (e.g. bacteria). Three approaches were tested: shotgun sequencing(3), targeting the whole human genome, and targeting ~850,000 custom SNPs. The two targeted techniques performed approximately the same, and both significantly outperformed the shotgun sequencing.
The whole-genome targeting approach, which is also used in Parabon’s law enforcement casework for highly degraded samples, was determined to be the most cost effective.
In traditional genome sequencing work, the goal is to sequence each piece of the human genome ~30 times, known as “30X coverage.” With this historical bone sample, even with targeting, each sequencing run only yielded ~2.5X coverage. Therefore, low-coverage imputation(4) was applied to create one SNP profile that could be used for genetic genealogy, kinship inference, ancestry, and phenotype prediction. Low-coverage imputation uses information from thousands of sequenced genomes to statistically infer the most likely genotype at each SNP, and it has been validated by Parabon for law enforcement casework(5).
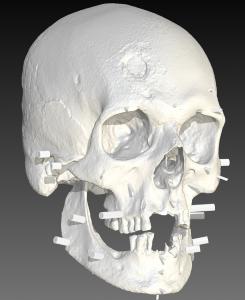
Using machine learning models built on published variants(6)(7), along with additional variants discovered by Parabon, JB55 was predicted to have Very Fair / Fair skin (92.2% confidence), Brown / Hazel eyes (99.8% confidence), Brown / Black hair (97.7% confidence), and Few / Some freckles (50.0% confidence). Using the phenotypic trait predictions and a digital 3D image of the skull, Thom Shaw, an IAI-certified forensic artist at Parabon, reconstructed JB55’s likely appearance.
Another individual in the cemetery was believed to be a relative of JB, so their DNA was also analyzed using whole-genome enrichment, sequencing, and low-coverage imputation. This sample was of much lower quality and only yielded ~0.68X coverage. Kinship analysis performed in Parabon’s Fx Forensic Analysis Platform(8) showed a 3rd-degree (1st cousin) relationship to JB. Both data files were successfully uploaded to the GEDmatch database, demonstrating that the high-quality genome-wide data needed for investigative genetic genealogy can be generated from highly degraded bones using these laboratory and bioinformatics protocols. Tracing the family trees of the GEDmatch matches led to ancestors with the surname Barber living in New England in the 18th and 19th centuries, supporting the hypothesis that his identity was most likely John Barber.
---
(1) Daniels-Higginbotham et al. (2019). DNA Testing Reveals the Putative Identity of JB55, a 19th Century Vampire Buried in Griswold, Connecticut. Genes, 10, 636. https://doi.org/10.3390/genes10090636
(2) Gorden et al. (2022). Extended kinship analysis of historical remains using SNP capture. Forensic Science International: Genetics, 57, 102636. https://doi.org/10.1016/j.fsigen.2021.102636
(3) Random sequencing of all DNA in the entire sample with no selection
(4) Hui et al. (2020). Evaluating genotype imputation pipeline for ultra low coverage ancient genomes. Scientific Reports, 10, 18542. https://doi.org/10.1038/s41598-020-75387-w
(5) Cady et al. (2022). Whole-genome sequencing of degraded DNA for investigative genetic genealogy. Forensic Science International: Genetics Supplement Series. https://doi.org/10.1016/j.fsigss.2022.09.008
(6) Chaitanya, et al. (2018). HIrisPlex-S system for eye, hair and skin colour prediction from DNA: Introduction and forensic developmental validation. Forensic Science International: Genetics, 35, 123–135. https://doi.org/10.1016/j.fsigen.2018.04.004
(7) Eriksson et al. (2010). Web-based, participant-driven studies yield novel genetic associations for common traits. PLoS Genetics, 6(6), 1–20. https://doi.org/10.1371/journal.pgen.1000993
(8) A commercially-available software package that reads NGS and WGS data from any sequencing instrument and performs both traditional and advanced STR and SNP analyses, such as identity matching, kinship inference, mixture deconvolution, and phenotype prediction.
https://parabon-nanolabs.com/dna-analytics/fx/
*About AFDIL: The Armed Forces Medical Examiner System’s Armed Forces DNA Identification Laboratory (AFDIL) is the Department of Defense’s only human remains DNA testing laboratory whose main missions are to support the Armed Forces Medical Examiner System, the Defense POW-MIA Accounting Agency and other Federal agencies with the DNA identification of all service members killed in current day operations, past conflict unknowns (World War II, Korea, Vietnam and the Cold War, and other unknown human remains. AFDIL leads the forensic community with developing new and novel forensic DNA testing methodologies to meet the needs or our fallen service members and their families.
Paula Armentrout
Parabon NanoLabs, Inc.
+1 703-689-9689 ext. 250
email us here
Legal Disclaimer:
EIN Presswire provides this news content "as is" without warranty of any kind. We do not accept any responsibility or liability for the accuracy, content, images, videos, licenses, completeness, legality, or reliability of the information contained in this article. If you have any complaints or copyright issues related to this article, kindly contact the author above.